User guide
Introduction
AgileAnnotator identifies sequences that have been aligned to the protein-coding sequences
or to regions within 50 bp of a splice junction. Each read is then screened for sequence mismatches and insertions or deletions
highlighted in its corresponding “CIGAR” data field in the SAM file. A position is called only if at least 5 reads
for the two most common alleles are mapped to the position. A position is called as heterozygous if 10% or more of the reads
identify the minor allele. The variant-calling algorithm is designed to be promiscuous, since the final cut-off parameters are
set interactively later, using AgileVariantViewer.
Table 1 illustrates the variant-calling process:
File formats
A description of the file formats for the variant and read depth files created by AgileAnnotator can be found
here.
Variant calling examples
| A | C | G | T | Read depth
(major+minor) | Minor allele percentage | Comments |
|---|
| 0 | 0 | 0 | 4 | 4 | 0% | No call, since read depth below 5 |
| 0 | 0 | 0 | 5 | 5 | 0% | Homozygous T |
| 1 | 0 | 0 | 4 | 5 | 20% | Heterozygous A/T, as minor allele is 20% of major+minor allele read depth |
| 1 | 1 | 0 | 3 | 4 | 25% | No call, as read depth of major plus a single minor allele is below 5 |
| 1 | 4 | 1 | 7 | 11 | 36% | No call, as minor allele read depth is not more than twice the combined read depth of the two remaining (presumably erroneous) bases (A+G) |
| 0 | 10 | 0 | 90 | 100 | 10% | Heterozygous A/T, since minor allele is 10% of major+minor allele read depth |
| 0 | 10 | 0 | 91 | 101 | 9.9% | Homozygous T, since minor allele is <10% of major+minor allele read depth |
| 0 | 10 | 0 | 89 | 99 | 10.1% | Heterozygous A/T, since minor allele is >10% of major+minor allele read depth |
| 0 | 10 | 4 | 90 | 100 | 10% | Heterozygous A/T, since minor allele is 10% of major+minor read depth |
Table 1
Creating an annotation file
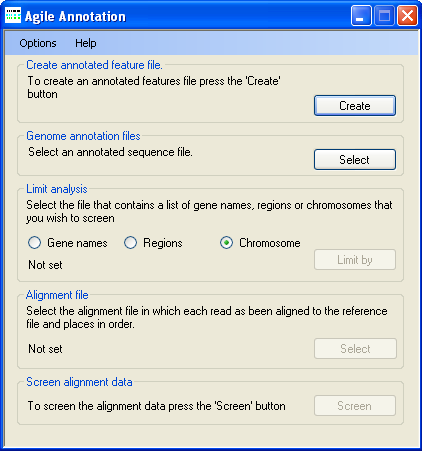
Figure 1: AgileVariantViewer user interface
AgileAnnotator is designed to identify sequence variants that can be suspected to disrupt a protein’ function.
It therefore only considers sequence variants that are within or close to protein-coding sequence.
To achieve this, AgileAnnotator makes use of an annotation file that contains the locations of protein-coding exons
and their genomic sequences. This file can be created either by AgileAnnotator or by
AgileGenotyper. To create an annotation file, first press the Create
button in the Create annotated feature file panel (Figure 1).
This causes the Annotation file creation window to be displayed (Figure 2).
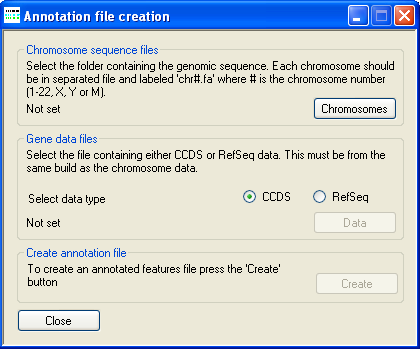
Figure 2: The annotation creation window
Note: The genomic sequences in the annotation files are derived from the current genomic sequence reference files and MUST match the version used
when aligning the sequence reads to the genome and they MUST also be the same build version used by the creators of the CCDS files, used to identify the
positions of coding sequences. If different reference sequences are used the analysis will fail!
The location of the uncompressed FASTA-format genomic reference files is selected using the Chromosomes button under
Chromosome sequence files. These reference sequence files must follow the specific naming convention where each file name starts with "chr"
followed by either the chromosome number or "X" or "Y", and has the .fa file extension. Permitted names include chr1.fa, chr5.fa for an autosome, while the X and Y may be named chrX.fa or chr23.fa and chrY.fa or chr24.fa, respectively.
Next press the CCDS button in the CCDS data files panel and select the file containing the positions
of the coding sequences as described by the Consensus CDS (CCDS) project. These files can be downloaded from the NCBI
CCDS web page or FTP site.
Finally, press the Create button under Create annotation file and enter a name for the genomic annotation
file. Since AgileAnnotator has to read all of the sequences in the genomic reference files and then write a large amount of data,
the creation of the annotation file may take several minutes.
Screening a SAM file for sequence variants
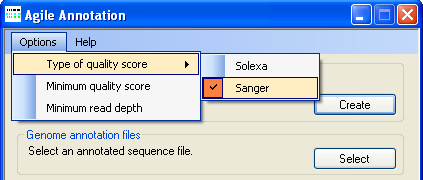
Figure 3: Adjusting the variant calling parameters
Before a SAM file is screened for sequence variants, it is necessary to select Solexa- or Sanger-type quality scores, using the
option, and and to adjust the variant calling cut-off parameters (quality and read depth), all
accessible under the menu (Figure 3).

Figure 4: Entering data
Once the variant calling options have been set, load the genome annotation file by pressing the Select button in the
Genome annotation file panel (Figure 4). Since this file is large, it may take a while to load. It is VERY important
that the annotation file was created using the same genomic reference sequence files used to align the sequence data in the SAM file.
It is possible to limit the exported sequence variants to those originating from a specified list of genes, genomic regions or chromosomes. To do this, select
the appropriate option in the Limit analysis panel and then press the Limit by button to choose a plain
text file containing the data. Figure 5 shows the format of these files, with X and Y indicating the sex chromosomes. Gene names must match those in
the CCDS file. To export variants from across the whole genome, create a chromosomal limits file containing
the numbers 1 to 24.
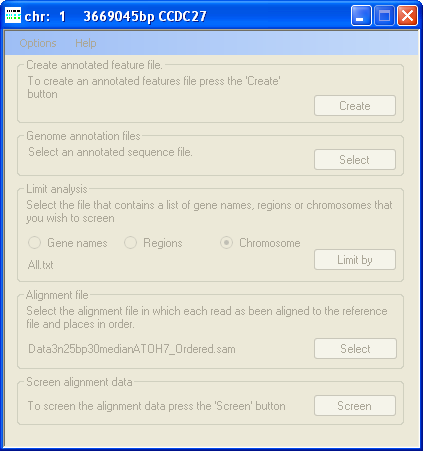
Figure 6: The title bar of AgileAnnotator shows the status of the analysis
Next, choose an ORDERED SAM file using the Select button in the Alignment file panel (Figure 4).
The aligned sequence reads in this file must be ordered by chromosome and chromosomal position.
Finally, press the Screen button in the Screen alignment data panel (Figure 6) and enter the name of the file to save the
exported sequence variants to. AgileAnnotator will then export the sequence variants as it reads the SAM file, showing the progress
of the analysis in the window title bar. AgileAnnotator will also generate a second file that contains read depth information
for each of the exons that sequence reads were mapped to. The latter file is designed to be used by AgileVariantViewer.
Since AgileAnnotator only stores the sequence reads for one gene at a time, it is does not require large amounts of memory, and the speed of the
analysis is limited by the speed at which it can read the SAM file, which should therefore be on a local hard drive. A description of the variant and read depth file formats
can be found here, while AgileVariantViewer and
AgileVariantViewer can be used to view the variant data.